COSC341
Theory of Computing
Alex Gavryushkin
![]()
2019 Semester 1
Table of contents
- Lecture 1: Introduction
- Lecture 2: Sets, relations, functions
- Lecture 3: Cardinality
- Lecture 4: Finite state automata
- Lecture 5: Non-deterministic automata
- Lecture 6: NFA = DFA
- Lecture 7: Pumping lemma
- Lecture 8: Pushdown automata and context free grammars
- Lecture 9: Regular languages
- Lecture 10: Classification of languages, pumping Lemma 2
- Lectures 11–13: Turing Machines
Handouts
Information
- COSC341 website
- Assessment
- Assignment 1: 10% due Thursday 28 March
- Assignment 2: 10% due Friday 10 May
- Assignment 3: 10% due Friday 24 May
- Final exam: 70% on TBA
Lecturer
Twitter: @bioDS_lab
Lecture 1
Introduction
Who cares about theory of computing?
I do!
ML = Machine Learning
CoSt = Computational Statistics
AS = Applied Statistics
Who cares about theory of computing?
You do!
Garey and Johnson. Computers and Intractability.
Why biological data science?
Because the skills are highly transferable:
- Noisy data
- Visualisation
- Communication
- High-performance computing
Why is it suddenly a thing?

Sets
A set is a collection of objects, completely determined by the objects. Two sets are equal if they contain the same objects.$X = \{1, 2, 5\} = \{2, 5, 1\} = \{5, 2, 1, 5, 1\}$
$X = \{x \mid P(x)\}$, where $P(x)$ is a logical condition on $x$.
$\mathbb N = \{0, 1, \ldots \}$
$X = \{x \mid (\exists y \in \mathbb N) x = y^2\} \stackrel{?}{=} \{x \mid x = y^2\}$
$9 \in X?$
Empty set
$\varnothing$
is a set!
You will forget this. Yes, you.Operations on sets
ML = Data $\cap$ Algorithms
(Statistics$\setminus$DS) $\cap$ Algorithms $\neq \varnothing$
$\overline{\mbox{Data} \cup \mbox{Algorithms} \cup \mbox{Statistics}} = ?$
Lecture 2
Sets, relations, functions
Subsets, ordered tuples, relations, and functions
$A \subseteq B \iff (\forall x) (x \in A \Rightarrow x \in B)$
$\mathcal P(A) = \{X \mid X \subseteq A\}$
$(a, b) = (c, d) \iff (a = c~\&~b = d)$
$A \times B = \{(a, b) \mid a \in A, b \in B\}$
$P \subseteq A \times B$ is a relation (from $A$ to $B$, or on $A$ if $A = B$)
$f \subseteq A \times B$ is a function if $(f(a) = b ~\&~ f(a) = c) \Rightarrow b = c$
Types of functions
Let $f : A \to B$. The function $f$ is called
injective if $f(a) = f(b) \Rightarrow a = b$
surjective if $(\forall y \in B)(\exists x \in A)f(x) = y$
bijective if $f$ is injective and surjective.
Molecular biology: crash course
Classical genetics
Modern genomics
Reality
Partitions and equivalence relations
$A = B \sqcup C$ is a partition if $A = B \cup C$ and $B \cap C = \varnothing$
A relation $\sim$ on $A$ is called an equivalence relations if $\sim$ is:
- Reflexive $(\forall x \in A)~x \sim x$
- Symmetric $(\forall x, y \in A)~(x \sim y \Rightarrow y \sim x)$
- Transitive $(\forall x, y, z \in A)~(x \sim y ~\&~ y \sim z \Rightarrow x \sim z)$
Theorem 1
A partition of a set $A$ is the same thing as an equivalence relation on $A$.In other words: Let $A$ be a set. Then
- A partition $A_1 \sqcup A_2 \sqcup \ldots$ of the set $A$ is defined by an equivalence relation on $A$.
- An equivalence relation $\sim$ on the set $A$ defines a partition on $A$.
Lecture 3
Cardinality
Set $A$ has smaller cardinality than $B$ if there exists a (total) injection $f : A \to B$, written $|A| \leqslant |B|$.
Theorem: $|A| = |B| \iff (|A| \leqslant |B| ~\&~ |B| \leqslant |A|)$.
$|\mathbb N| \stackrel{?}{=} |\mathbb Z| \stackrel{?}{=} |\mathbb Q| \stackrel{?}{=} |\mathbb R|$
Theorem 2
$|\mathbb N| < |\mathbb R|$Lecture 4
Finite state automata
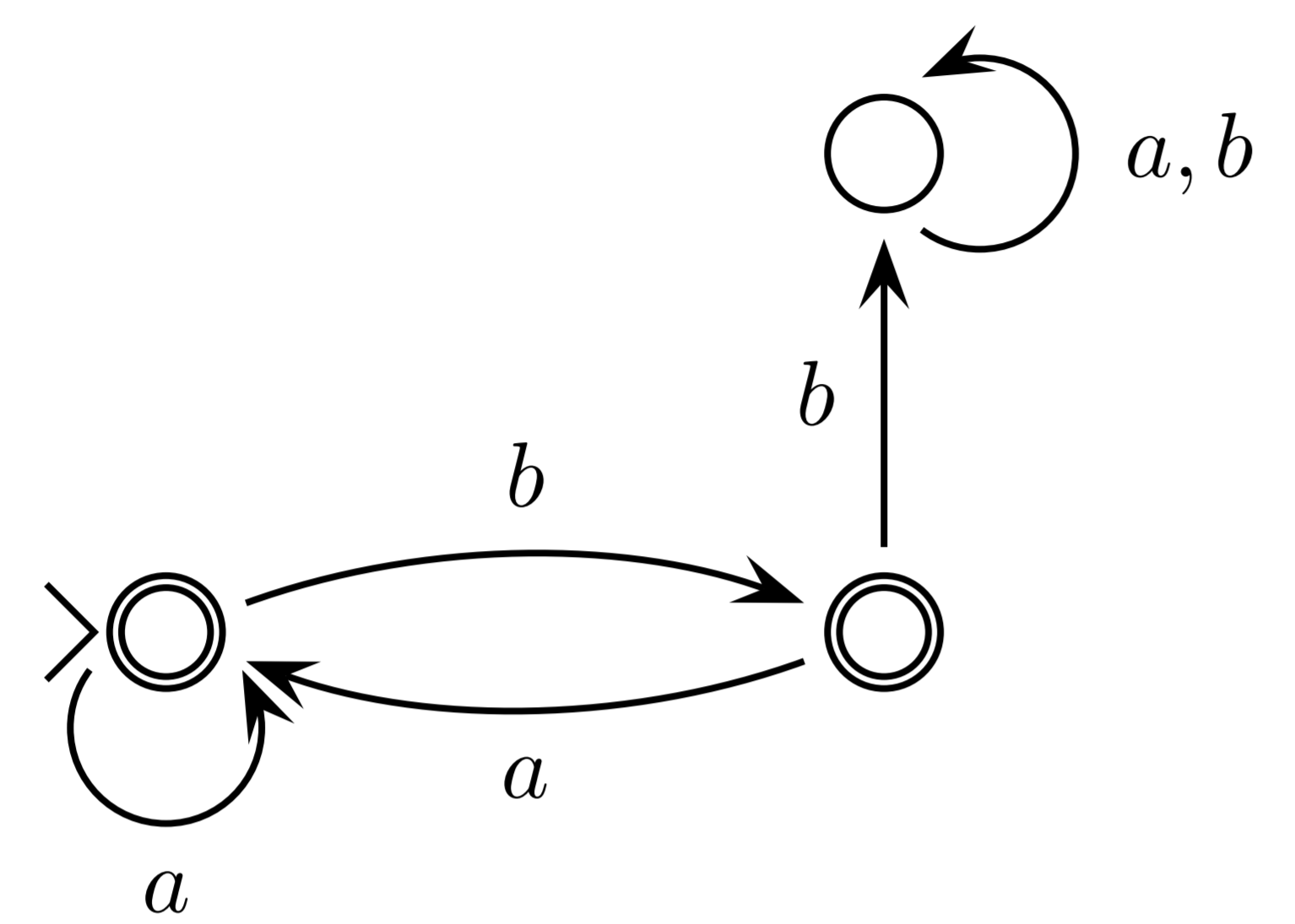
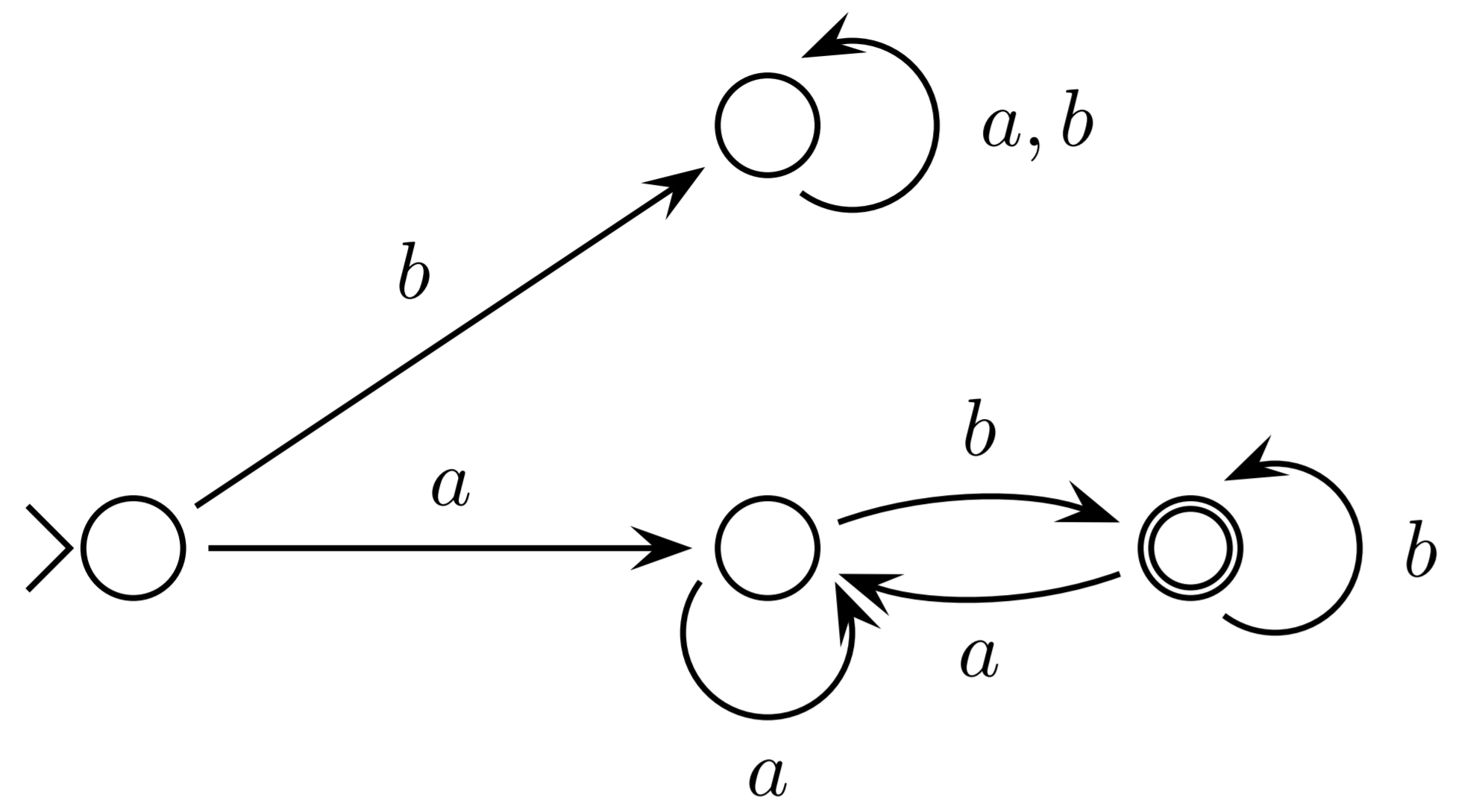
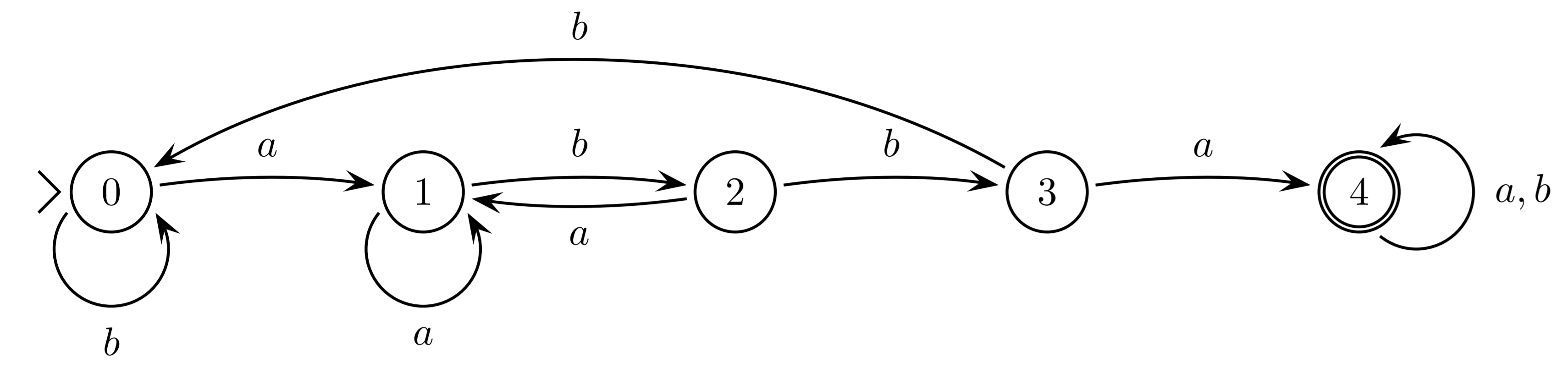
DFA

A deterministic finite state automaton, $M$, consists of:
- A finite set, $Q$, of states
- A finite set $\Sigma$ called the alphabet
- A total function $\delta: Q \times \Sigma \to Q$ called the transition function
- A distinguished state $q_0 \in Q$ called the initial state
- A subset $F \subseteq Q$ called the final or accepting states.

Given a word $w = w_0 w_1 \dots w_{n-1} \in \Sigma^*$ the computation carried out by $M$ on input $w$ is a sequence of states $q_0, q_1, q_2, \dots, q_n$ defined as follows: \[ q_1 = \delta(q_0, w_0), \: q_2 = \delta(q_1, w_1), \: \dots ,\: q_n = \delta(q_{n-1}, w_{n-1}) \]
We say that $M$ accepts or recognises $w$ if $q_n \in F$ and otherwise it rejects $w$.
The language of $M$, $L(M)$ is just the set of strings in $\Sigma^*$ that $M$ accepts.
Lecture 5
Non-deterministic automata
NFA
Relax the definition of the transition function to become a transition relation. \[ \delta \subseteq Q \times \Sigma \cup \{\lambda\} \times Q \]
$\{ab\}$ in English alphabet
Examples
Design an automaton that recognises the following languages in alphabet $\{a,b\}$
$L_1 = \{w | w = xaaybbbz, \mbox{ where } x,y,z \in\{a,b\}^*\}$
$L_2 = \{w | w \mbox{ contains $aa$ or $bbb$}\}$
$L_3 = \{w | w \mbox{ contains $aa$ and $bbb$}\}$
Lecture 6
NFA = DFA, pumping lemma
Theorem 3
The classes of languages recognised by DFA and NFA coincide.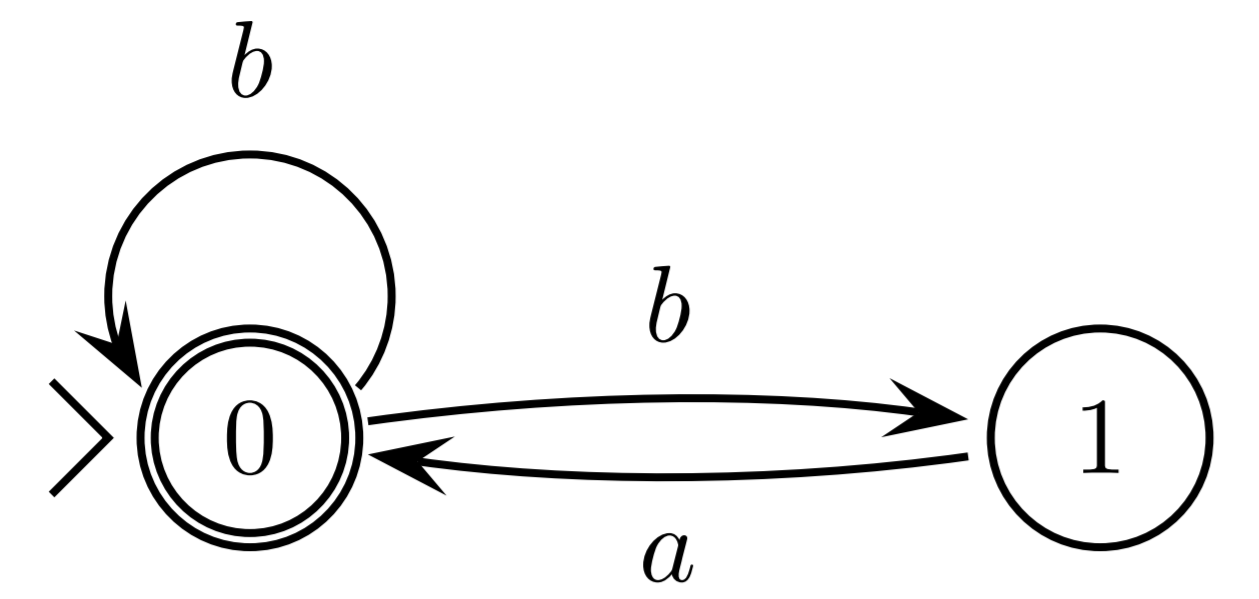
Theorem 3
The classes of languages recognised by DFA and NFA coincide.Proof: Go to the lecture (or read in the notes).
Lecture 7
Pumping lemma
Pumping lemma
Let $L$ be an automatic language.
Then there exists a positive integer $k$ such that if $z \in L$, $|z| \geq k$ then for some $u$, $v$, and $w$:
$
\begin{eqnarray*}
z &=& u \cdot v \cdot w \\
|u| + |v| &\leq& k \\
|v| &>& 0 \\
uv^iw &\in& L \quad \mbox{for all $i \geq 0$}.
\end{eqnarray*}
$
Proof: Haven't we just proven it?
$L = \{w \in \{a, b\} \mid \mbox{$w$ contains an odd number of $a$'s}$
$\mbox{and an even number of $b$'s}\}$
Lecture 8
Pushdown automata and context free grammars
- A finite set $Q$ of states
- A finite set $\Sigma$ called the input alphabet (lower case letters)
- A finite set $\color{blue}{\Gamma}$ called the stack alphabet (upper case letters)
- A relation $\delta \subseteq Q \times \Sigma \cup \{\lambda\} \times \color{blue}{\Gamma \cup \{\lambda\}} \times Q \times \color{blue}{\Gamma \cup \{\lambda\}}$
called the transition relation - A distinguished state $q_0 \in Q$ called the initial state
- A subset $F \subseteq Q$ called the final or accepting states.
The stack should be empty to accept a string!
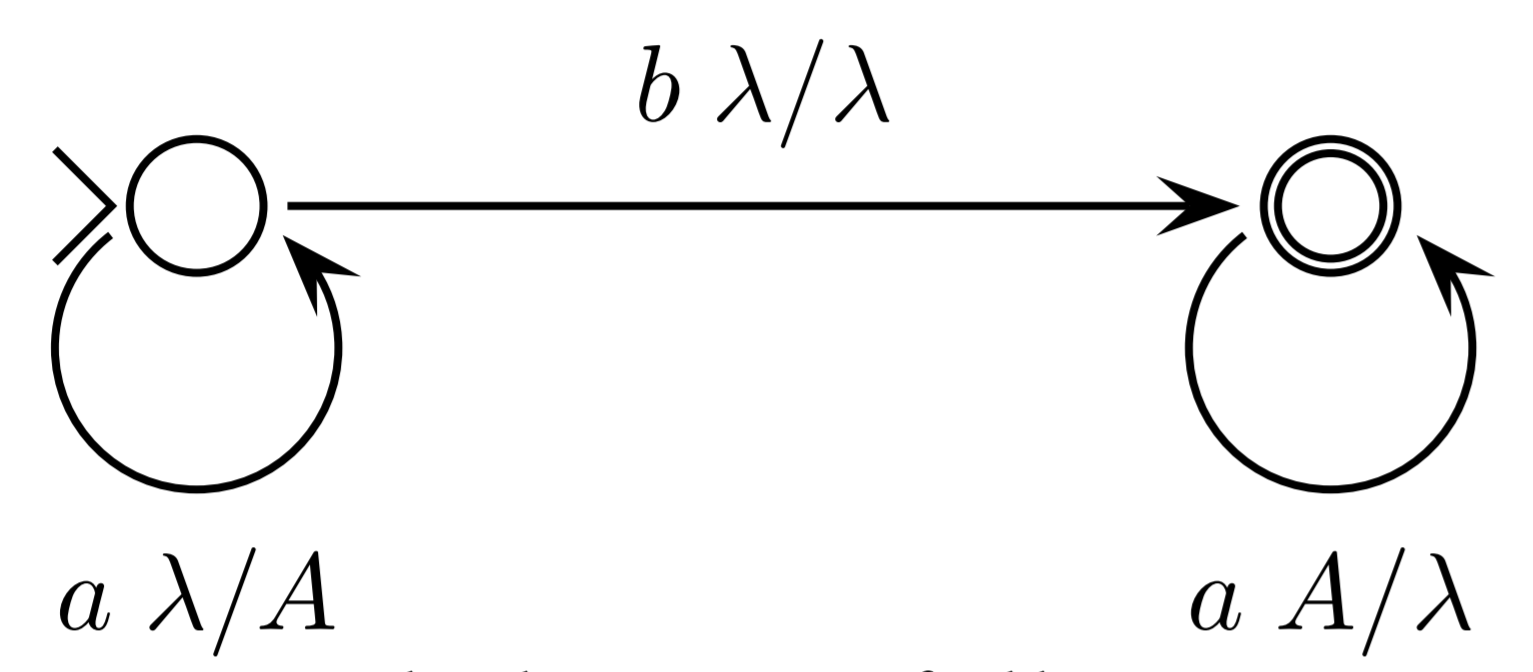
- A finite set $V$ of nonterminals (or variables)
(upper case letters) - A finite set $\Sigma$ called the alphabet (or terminals)
(lower case letters), $\Sigma \cap V = \varnothing$ - A finite set $P$ of production rules,
which is a subset of $V \times (V \cup \Sigma)^*$ - A distinguished $S \in V$ called the start symbol
Examples
$S \to aS, S \to bS, S \to \lambda$
$S \to aS, S \to bT, T \to bT, T \to \lambda$
$\{a^nb^n \mid n \in \mathbb{N}\}$
Theorem
$\mathcal L_{PDA} = \mathcal L_{CFG}$Genome phasing
Schwarz, Roland F., Anne Trinh, Botond Sipos, James D. Brenton, Nick Goldman, and Florian Markowetz. 2014. “Phylogenetic Quantification of Intra-Tumour Heterogeneity.” PLoS Computational Biology 10 (4): e1003535.
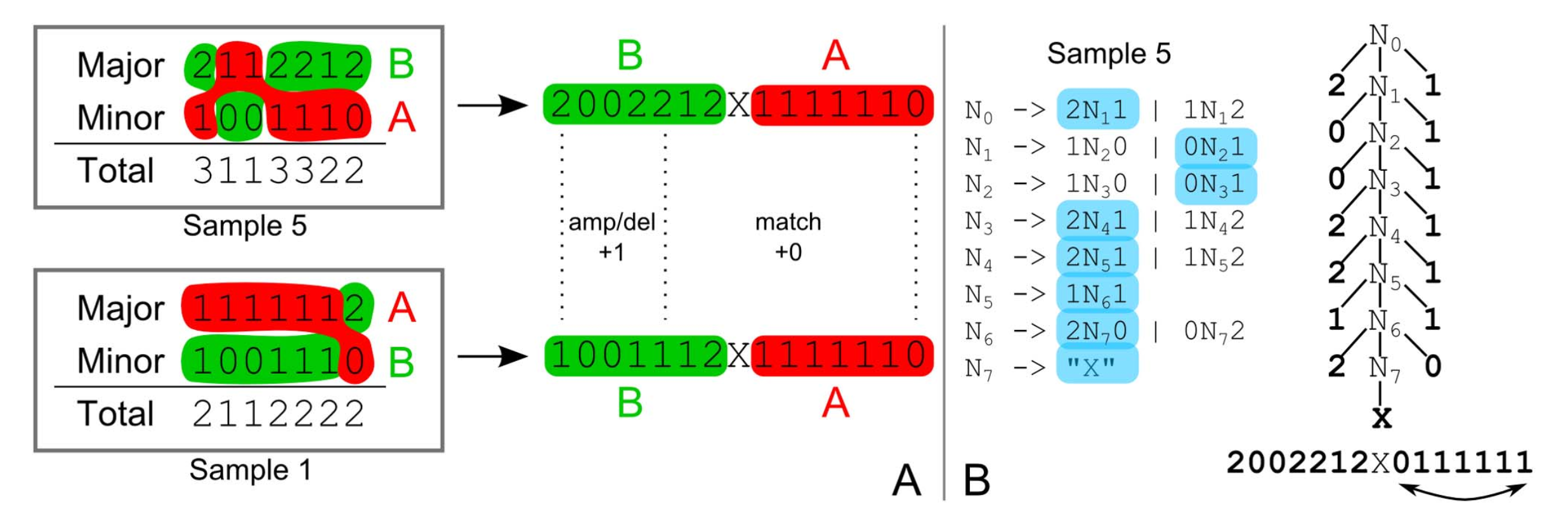
Schwarz, Roland F., Anne Trinh, Botond Sipos, James D. Brenton, Nick Goldman, and Florian Markowetz. 2014. “Phylogenetic Quantification of Intra-Tumour Heterogeneity.” PLoS Computational Biology 10 (4): e1003535.
Lecture 9
Regular languages
$\varnothing$; $\lambda$; $x$ for every $x \in \Sigma$.
If $\alpha$ and $\beta$ are regular expressions then so are the following:
$(\alpha\beta)$; $(\alpha \cup \beta)$; $\alpha^*$
The language $L_\alpha$ described by the regular expression $\alpha$ is defined in the following way:
$L_\alpha = \varnothing$ if $\alpha = \varnothing$
$L_\alpha = \{\lambda\}$ if $\alpha = \lambda$
$L_\alpha = \{x\}$ if $\alpha = x$
$L_{\alpha\beta} = \{uv \mid u \in L_\alpha \mbox{ and } v \in L_\beta\}$
$L_{\alpha\cup\beta} = L_\alpha \cup L_\beta$
$L_{\alpha^*} = \{w_1 \ldots w_k \mid w_1, \ldots, w_k \in L_\alpha \mbox{ and } k \in \mathbb N\}$
Language $L$ is called regular if it can be described by a regular expression $\alpha$, that is, there exists $\alpha$ such that $L = L_\alpha$.
Examples
\begin{align*} L_a &= a(a \cup b)^*\\ \\ L_e &= ((a \cup b)(a \cup b))^*\\ \\ L_{bb} &= (a \cup b)^*bb(a\cup b)^*\\ \\ L_{2b} &= a^*ba^*ba^* \end{align*}Lecture 10
Classification of languages, Pumping Lemma 2
Proof: Easy.
Theorem (ii): Automatic languages are closed under
$\cdot$ (concatenation), $\cup$ (union), $\cap$ (intersection),
and $\overline{\phantom a}$ complement.
Proof: Even easier.
Theorem (iii): Context-free languages are closed under
$\cdot$ (concatenation) and $\cup$ (union),
but not $\cap$ (intersection) or $\overline{\phantom a}$ complement.
Proof: Not yet possible — we need a version of the pumping lemma.
Classification of languages
(so far)$\mathcal L_{DFA} = \mathcal L_{NFA} = \mathcal L_{reg} \subset \mathcal L_{PDA} = \mathcal L_{CFG}$
Pumping Lemma 2
Let $L$ be a context free language. There is a positive integer $k$ such that for all $z \in L$ with $|z| \geq k$ we can write $z = uvwxy$ such that: \[ \begin{array}{c} |vwx| \leq k \\ |v| + |x| > 0 \\ uv^iwx^iy \in L \quad \mbox{for all $i \geq 0$}. \end{array} \]$\cdot$ (concatenation) and $\cup$ (union),
but not $\cap$ (intersection) or $\overline{\phantom a}$ complement.
Proof: Super easy now.
Note: However, if $L$ is a context-free language and $R$ a regular language then $L \cap R$ is context-free!
Lectures 11–13
Turing Machines
Turing machine
- A finite set $\Gamma$ called the alphabet (or tape symbols)
- A finite set $Q$ of states
- A distinguished state $q_0 \in Q$ called the starting state
- A subset $F \subseteq Q$ of halting states
- A distinguished symbol $\beta \in \Gamma$ called the blank symbol (or the empty cell)
- A partial function $F: (Q \setminus F) \times \Gamma \to Q \times \Gamma \times \{L, R, \varnothing\}$
called the programme
Turing machines can be used to formalise all kinds of algorithmic problems, including language recognition, generation, etc.
The program does not terminate otherwise.
Hence a non-terminating programme either hangs (infinite loop) or crashes (reaches a state and tape symbol with no command to execute).
We will be starting Turing machines on $$ \beta\underset{\Delta}{a_1} a_2 \ldots a_n \beta $$
$$ \begin{align} \mbox{Example 2} &\\ & q_0 \beta \to \mathrm{accept} \beta\\ & q_0 0 \to q_0 \beta R\\ & q_0 1 \to q_0 \beta R \end{align} $$
$$ \begin{align} \mbox{Example 3} &\\ & q_0 \beta \to \mathrm{accept} \beta\\ & q_0 0 \to q_0 1 R\\ & q_0 1 \to q_0 0 R \end{align} $$
for all $x$ and $y$: $$ f(x) = y $$ $$ \Updownarrow $$ $$ \beta \underbrace{1 \ldots 1}_{x \mbox{ times}} \beta \Rightarrow_M \beta \underbrace{1 \ldots 1}_{y \mbox{ times}} \beta \mbox{ and $M$ halts} $$
We denote the latter by $M(x) = y$.
We write $M(x)\downarrow$ for $M$ halts on $\beta \underbrace{1 \ldots 1}_{x \mbox{ times}} \beta$
for all $x_1, x_2$, and $y$: $$ f(x_1, x_2) = y $$ $$ \Updownarrow $$ $$ \beta \underbrace{1 \ldots 1}_{x_1 \mbox{ times}} 0 \underbrace{1 \ldots 1}_{x_2 \mbox{ times}} \beta \Rightarrow_M \beta \underbrace{1 \ldots 1}_{y \mbox{ times}} \beta \mbox{ and $M$ halts} $$ Similarly for more than two variables $x_1, x_2, \ldots, x_n$.
Example 2: implement the following module COPY. $$ \beta \underbrace{1 \ldots 1}_{x \mbox{ times}} \beta \Rightarrow_M \beta \underbrace{1 \ldots 1}_{x \mbox{ times}} 0 \underbrace{1 \ldots 1}_{x \mbox{ times}} \beta $$
Example 3: compute: $$ f(x, y) = xy $$
$a^* b^*$
$\{a^n b^n \mid n \in \mathbb N\}$
Universal Turing Machine
Theorem: There exists a Turing Machine $U$ which computes all computable functions.That is, for every computable function $f(x)$ there exists a number $s$ such that $U(s, x) = y$ if and only if $f(x) = y$.
Halting problem
Theorem: Not everything is computable.For example, the following function is not computable $$ h(x) = \begin{cases} U(x, x) + 1, & \mbox{ if } U(x, x) \downarrow\\ 0, & \mbox{ otherwise} \end{cases} $$
